
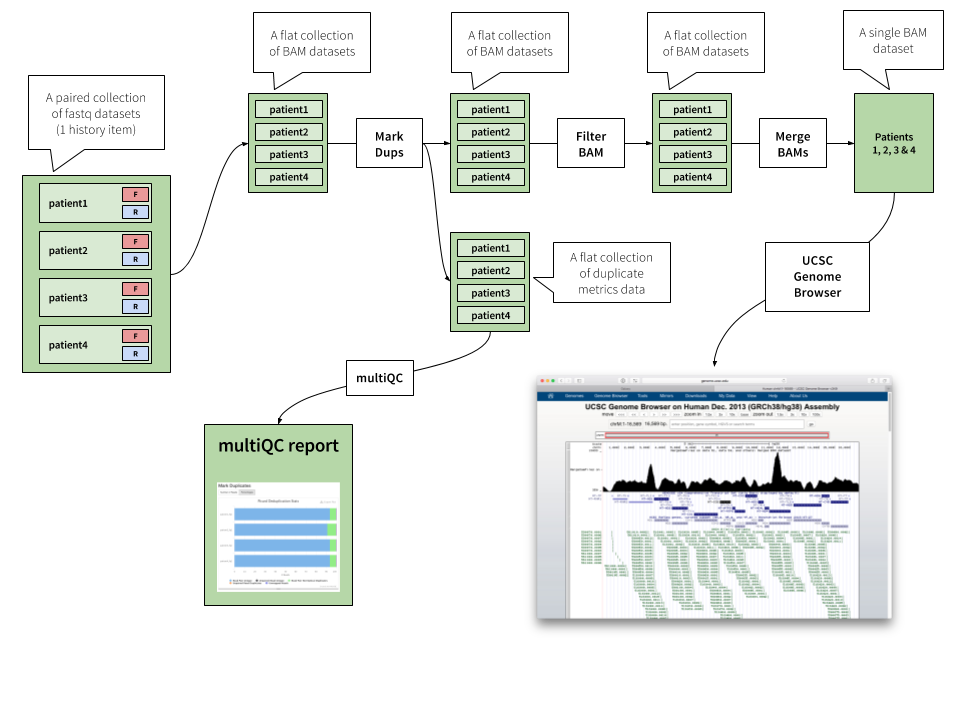
Flow chart for data analysis. This example shows a workflow for the... | Download Scientific Diagram
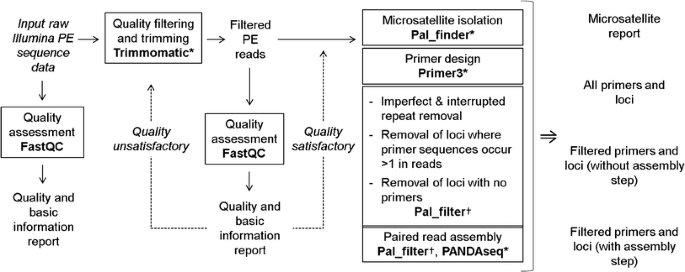
A Galaxy-based bioinformatics pipeline for optimised, streamlined microsatellite development from Illumina next-generation sequencing data | SpringerLink

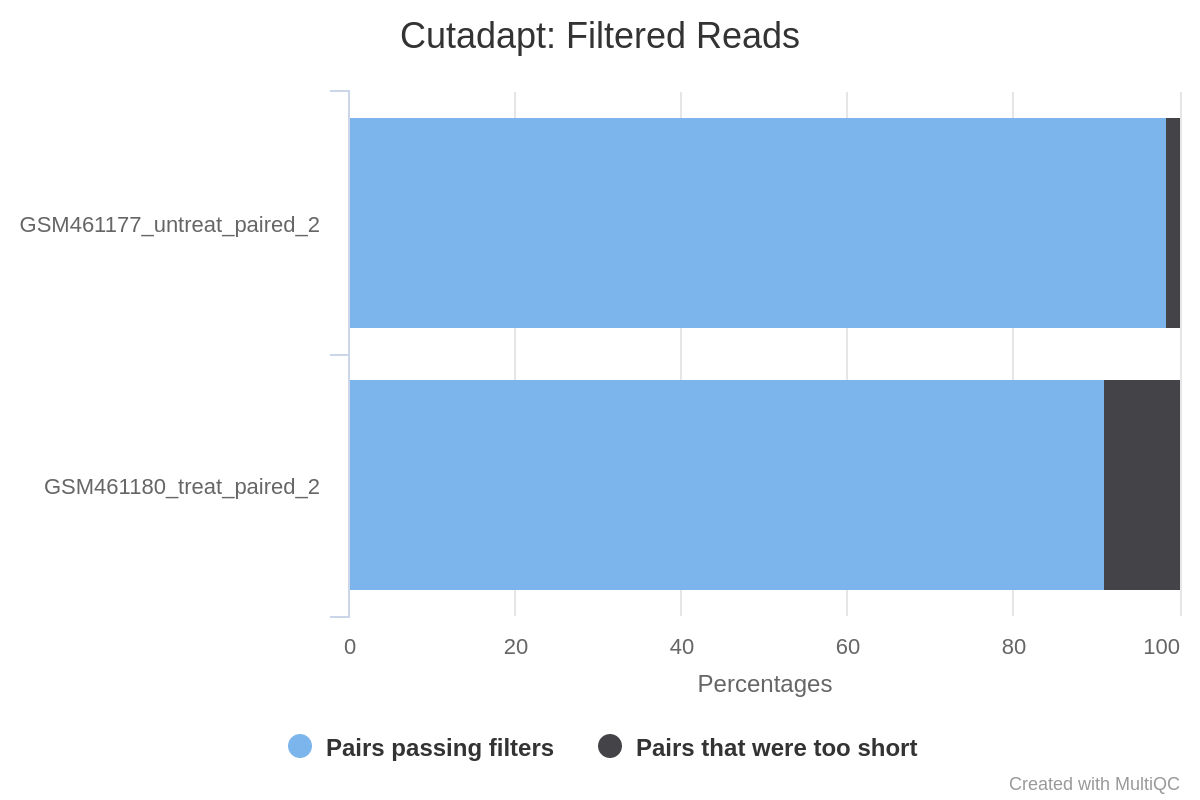
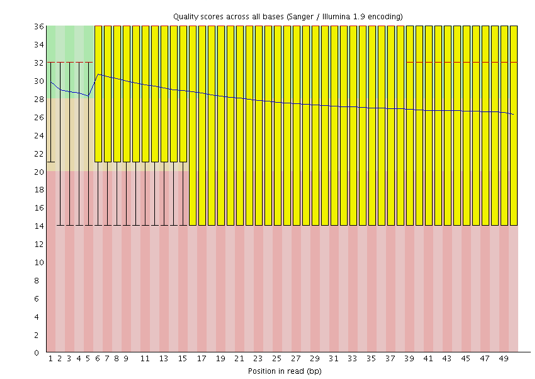
![Tutorial] Trailing of paired end reads using Trimmomatic tool in GALAXY. — Bioinformatics Review Tutorial] Trailing of paired end reads using Trimmomatic tool in GALAXY. — Bioinformatics Review](https://bioinformaticsreview.com/wp-content/uploads/2021/10/trimmomatic.jpg)

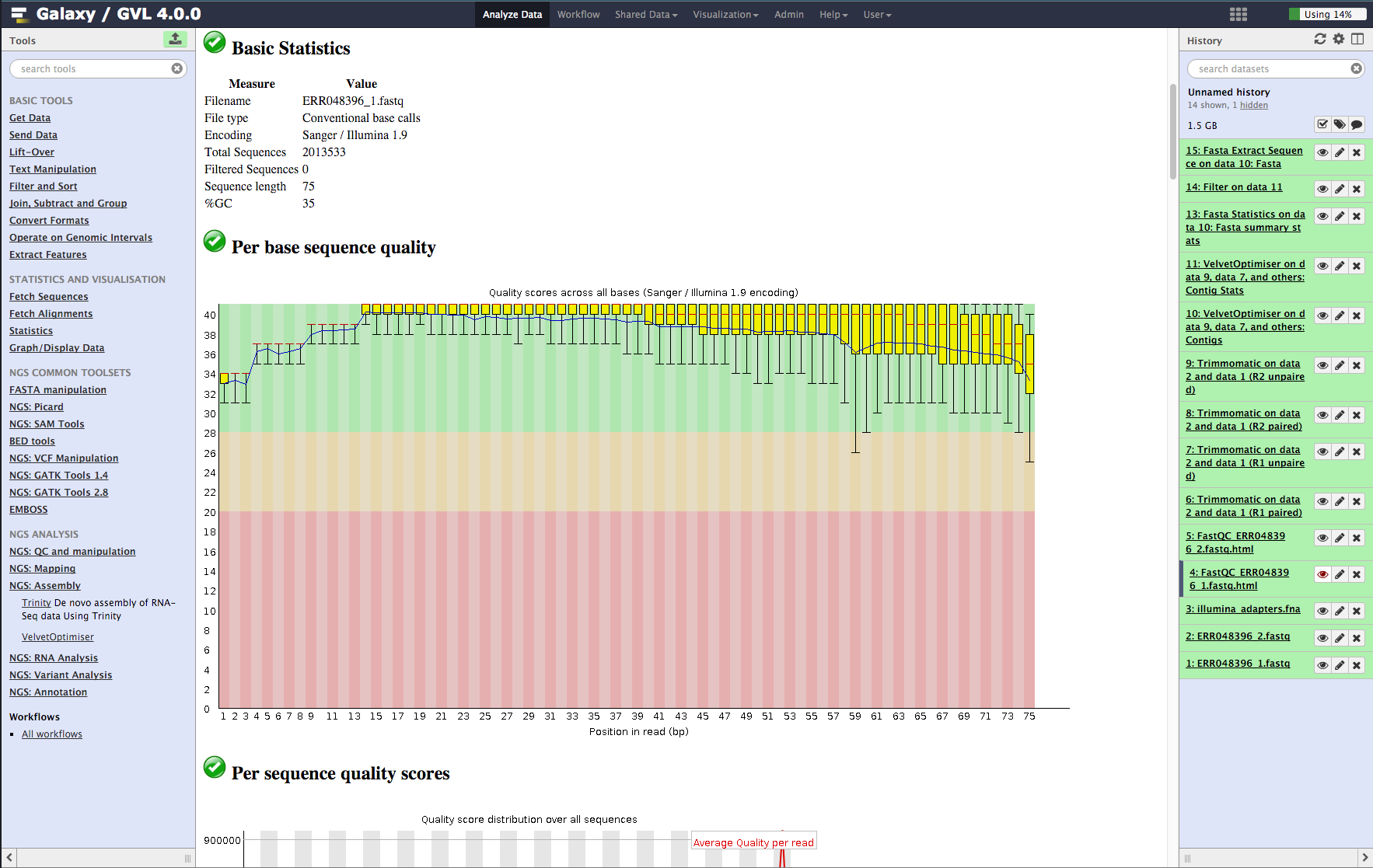


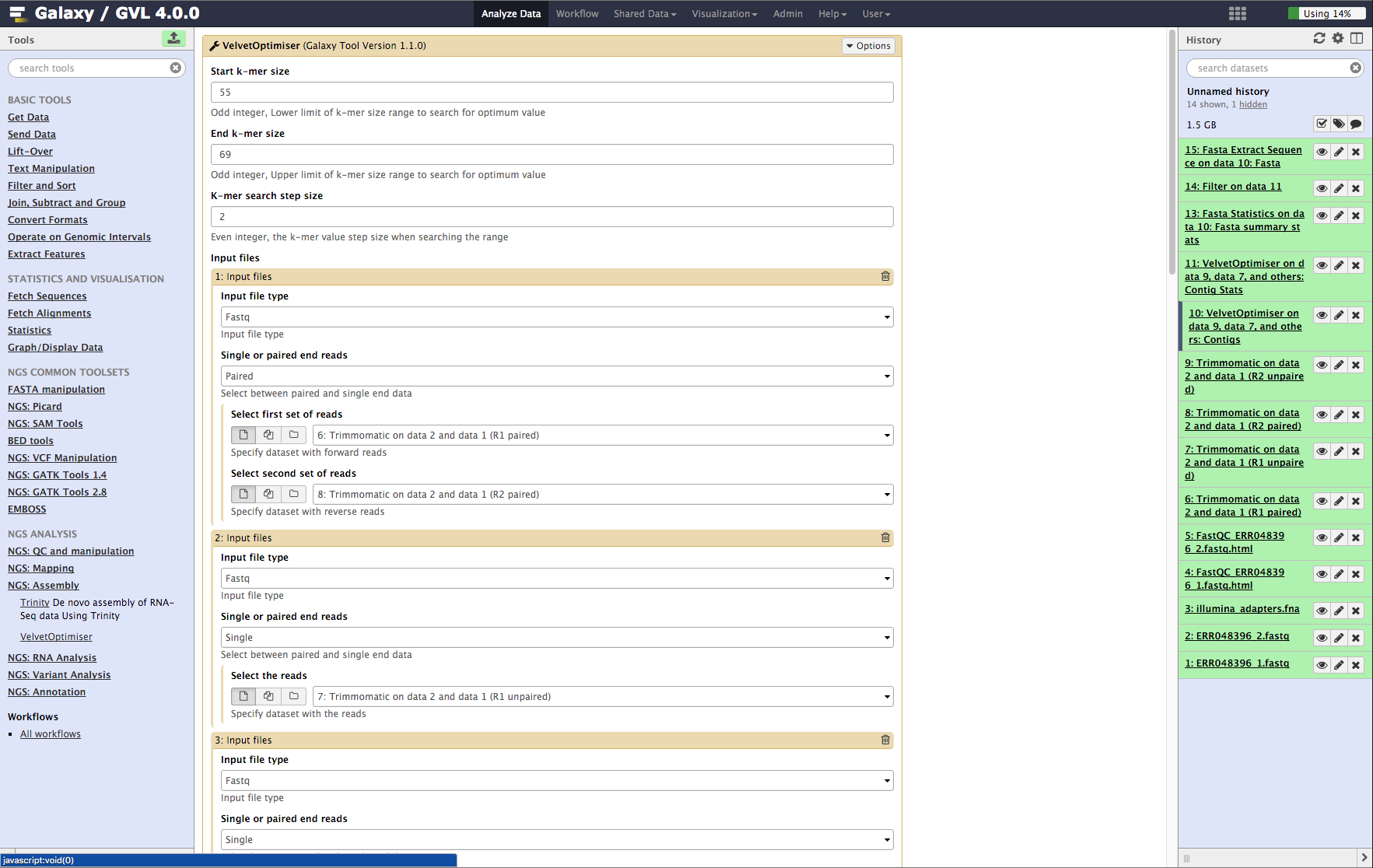





![Tutorial] Trailing of paired end reads using Trimmomatic tool in GALAXY. — Bioinformatics Review Tutorial] Trailing of paired end reads using Trimmomatic tool in GALAXY. — Bioinformatics Review](https://bioinformaticsreview.com/wp-content/uploads/2020/09/galaxy.jpg)