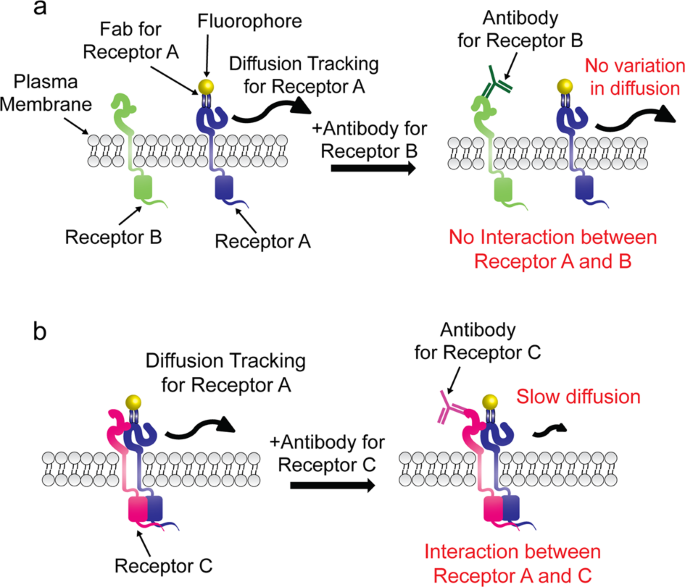
Analysis of transient membrane protein interactions by single-molecule diffusional mobility shift assay | Experimental & Molecular Medicine

Predicting transmembrane protein topology with a hidden markov model: application to complete genomes - ScienceDirect

Improved sequence-based prediction of interaction sites in α-helical transmembrane proteins by deep learning - ScienceDirect
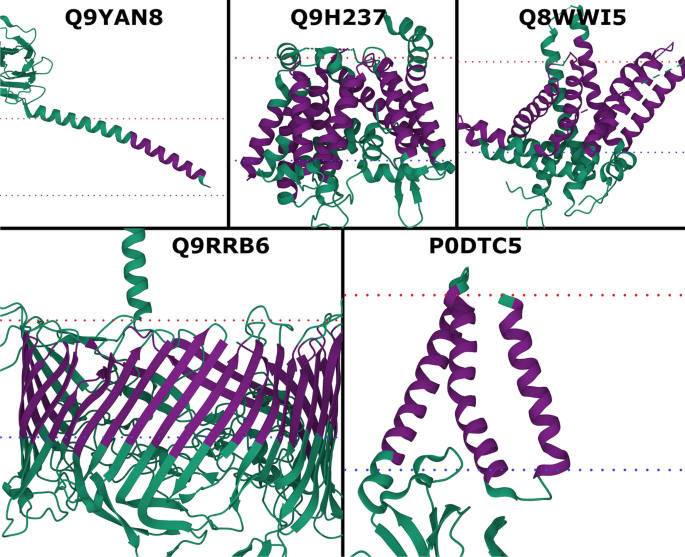
TMbed: transmembrane proteins predicted through language model embeddings | BMC Bioinformatics | Full Text
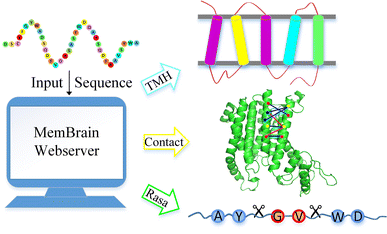
MemBrain: An Easy-to-Use Online Webserver for Transmembrane Protein Structure Prediction | Nano-Micro Letters

Methods for Systematic Identification of Membrane Proteins for Specific Capture of Cancer-Derived Extracellular Vesicles - ScienceDirect
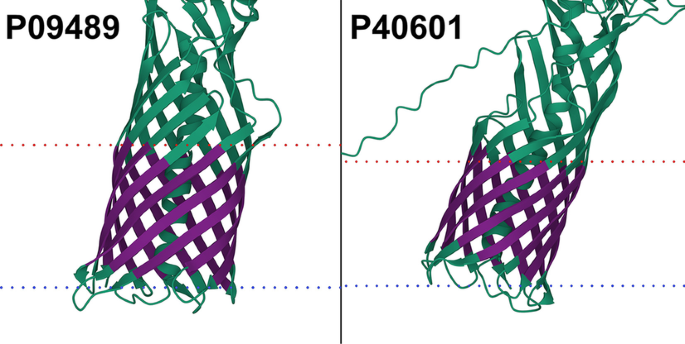
TMbed: transmembrane proteins predicted through language model embeddings | BMC Bioinformatics | Full Text
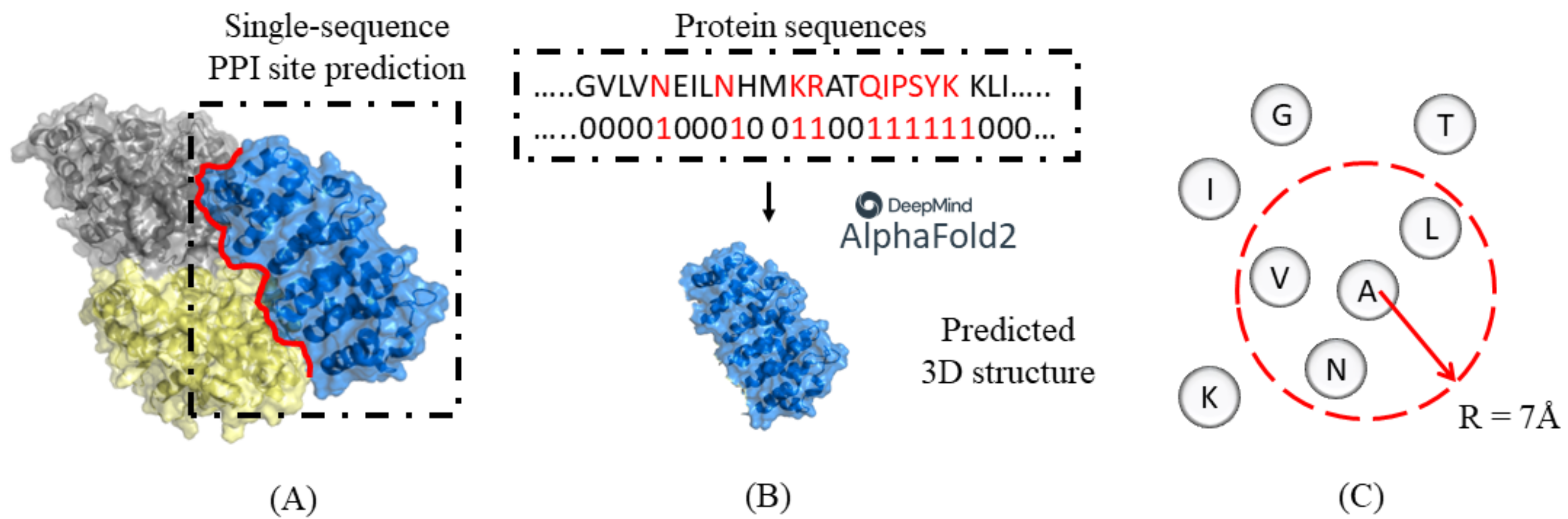
Biology | Free Full-Text | Evaluation of the Effectiveness of Derived Features of AlphaFold2 on Single-Sequence Protein Binding Site Prediction

The graphical output of TMHMM, showing the posterior probabilities for... | Download Scientific Diagram
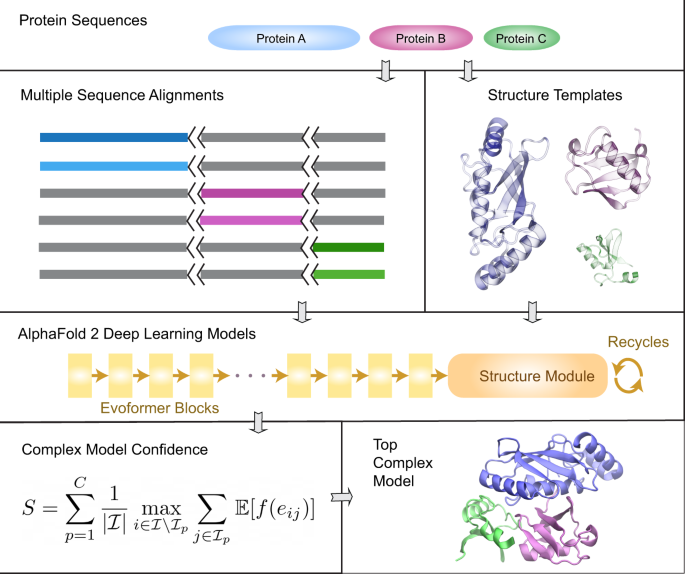
AF2Complex predicts direct physical interactions in multimeric proteins with deep learning | Nature Communications

IJMS | Free Full-Text | RLPredictiOme, a Machine Learning-Derived Method for High-Throughput Prediction of Plant Receptor-like Proteins, Reveals Novel Classes of Transmembrane Receptors

Accurate de novo structure prediction of large transmembrane protein domains using fragment-assembly and correlated mutation analysis | PNAS

Revealing the mechanisms of membrane protein export by virulence-associated bacterial secretion systems | Nature Communications
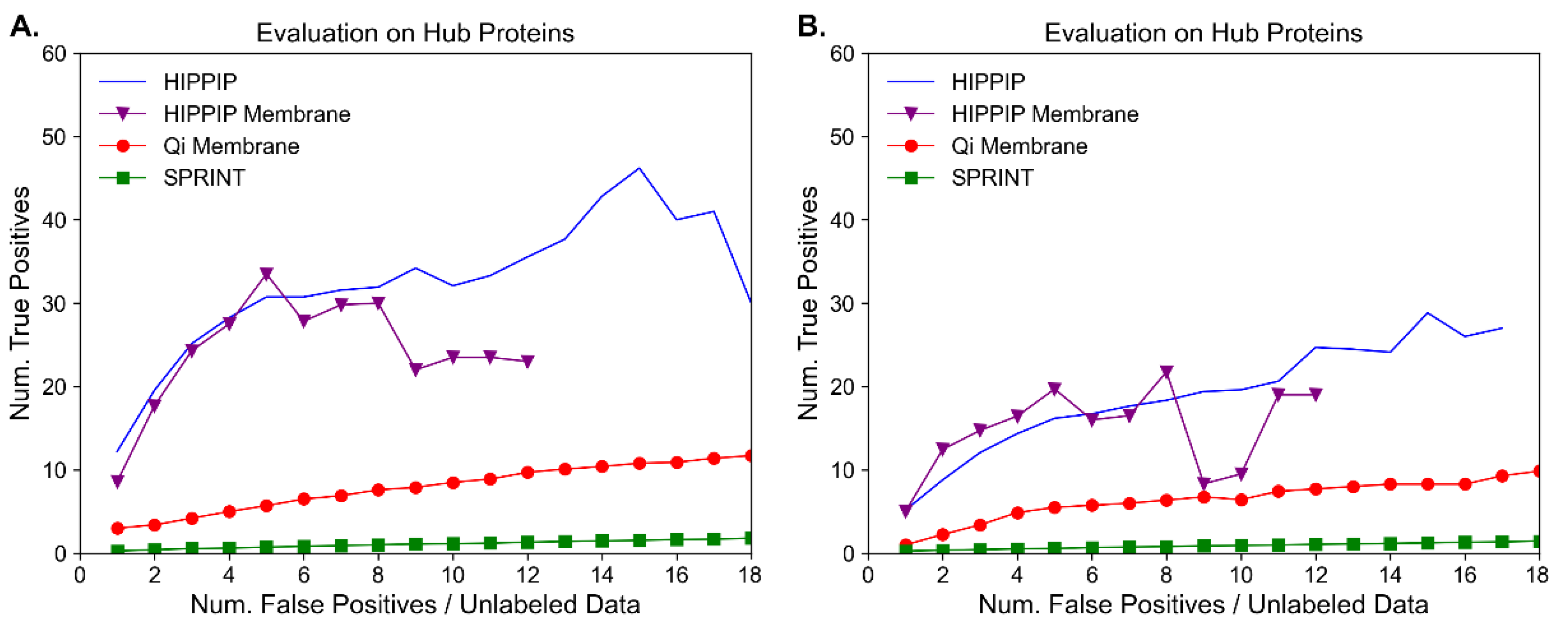
Molecules | Free Full-Text | Benchmark Evaluation of Protein–Protein Interaction Prediction Algorithms

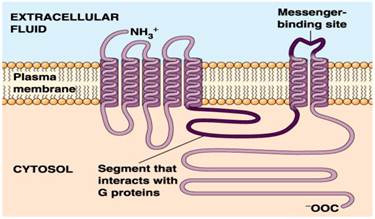




![RFPDR: a random forest approach for plant disease resistance protein prediction [PeerJ] RFPDR: a random forest approach for plant disease resistance protein prediction [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2022/11683/1/fig-2-full.png)